| Concept BIOISIS is an open access database dedicated to the study of biological macromolecules by small angle X-ray scattering (SAXS). The project is supported by the Department of Energy Office of Science Integrated Diffraction Analysis Technologies, the National Cancer Institute Structural Cell Biology of DNA Repair Machines and the National Institute of General Medical Sciences project MINOS (Macromolecular INsights Optimized by Scattering). BIOISIS aims to become the complete source for the deposition, distribution and maintenance of small angle X-ray scattering data and technologies. |
| Design BIOISIS was built with the the Ruby on Rails framework (Ruby 1.9.3 and Rails 3.2.8) and the MySQL relational database. Its chief designer and architect is Robert P. Rambo, Ph. D., of the Lawrence Berkeley National Lab. The need for a SAXS database for biological sciences was pursued by John Tainer, Ph.D., of the Scripps Research Institute in La Jolla, CA., nd Greg Hura, Ph.D., of the Lawrence Berkeley National Lab in Berkeley, CA. The database is designed around the concept of an “experiment” and relates a specific experiment to a set of genes, organisms, computational models and experimental data. 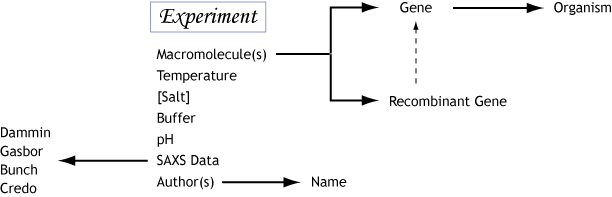 If we consider the fact that the structure of a macromolecule in solution is condition specific, then the data and models from a refined SAXS experiment will reflect a condition specific conformational state. Since SAXS experiments at synchrotron sources are highly efficient, a typical data collection at a synchrotron may sample several different experimental conditions, thus yielding measurements of different conformational states. This was an important realization incorporated into the design of BIOISIS. In contrast, those structures obtained from X-ray crystallography often measure a single conformational state due to both the nature of crystallography and the use of cryogenic temperatures during the experiment. |